HDAC9 Target Analysis Report Summary


About the Target
Based on the context information provided, HDAC9 is a histone deacetylase enzyme that regulates various cellular processes. HDAC inhibitors can increase the acetylation of histone and non-histone proteins, leading to both epigenetic and non-epigenetic effects on cell function [1]. In the context of Treg cells, HDAC9 plays a role in inhibiting FOXP3 transcription, which is necessary for Treg activation. Increased expression of HDAC9 in Treg cells may suppress their activation and lead to expansion of inflammatory cells [2]. HDAC activity is also involved in the regulation of HIF (hypoxia-inducible factor) transcription factors. HDAC inhibitors can block HDAC activity, destabilizing HIF1alpha and affecting the activation of hypoxia response genes [3]. Additionally, HDACs can interact with BET proteins and affect the presence of BRD4 on chromatin, ultimately altering transcriptional regulation [4]. HDAC inhibitors can also reduce HIF signaling and CXCR4 expression, potentially serving as therapeutics for inhibiting metastasis [5]. Overall, HDAC9 plays a role in regulating gene transcription, cellular function, and mediating the effects of hypoxia on gene expression.
Based on the provided context information, the following key viewpoints can be summarized regarding HDAC:
Butyrate, a compound primarily utilized as an energy source in normal colonocytes, can accumulate in the nucleus of cancerous colonocytes where it acts as an HDAC inhibitor to regulate genes involved in cell proliferation and apoptosis [6].
Inhibition of HDACs can lead to abnormal increase in Bmp acetylation and expression, affecting the formation and branching morphogenesis of the prostate [7].
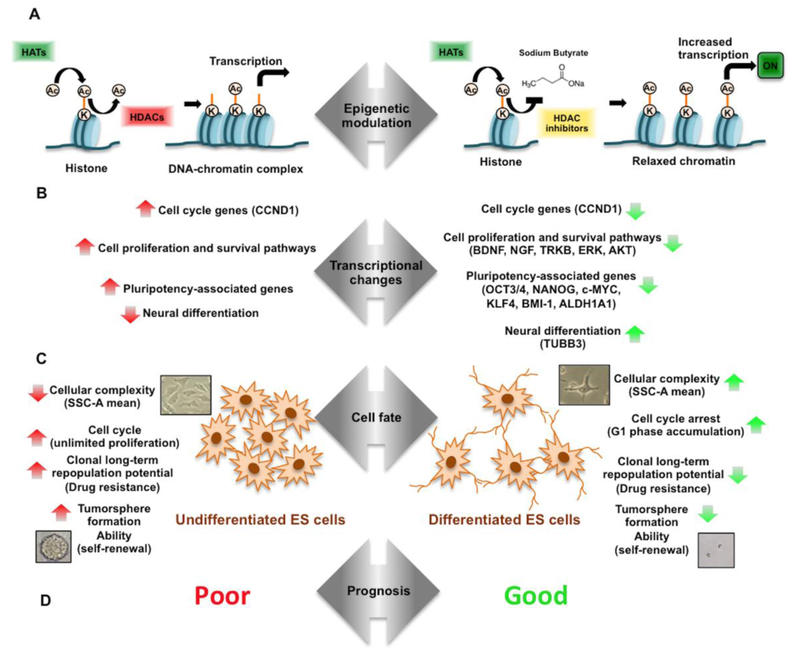
HDACs play a role in histone acetylation, which can control transcriptional inactivation and activation of genes [8].
The Set2-Rpd3S HDAC pathway is important for maintaining optimal expression dynamics of mRNAs and non-coding RNAs, with histone deacetylation causing a delay in the induction of target mRNA genes [9].
HDAC inhibitors can be employed to revert the aberrant epigenetic state often found in EWS cells, leading to changes in gene expression and promoting cellular differentiation [10].
Please note that the references to the corresponding context information are indicated with numbers in the summary for citation purposes.
Figure [1]

Figure [2]
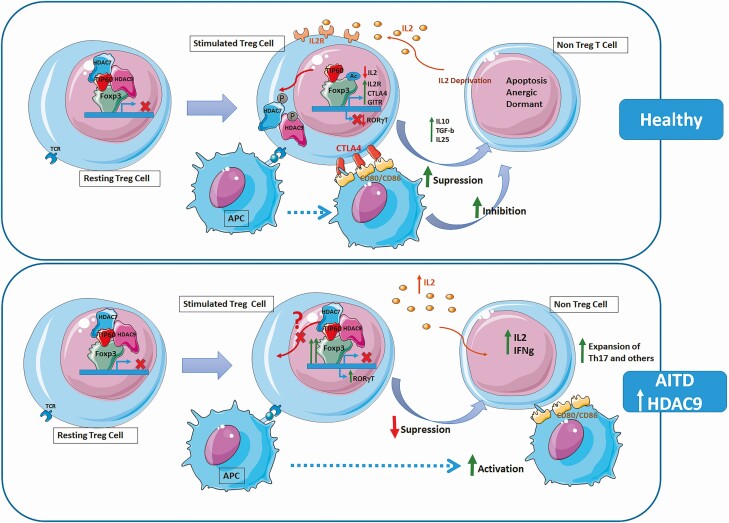
Figure [3]
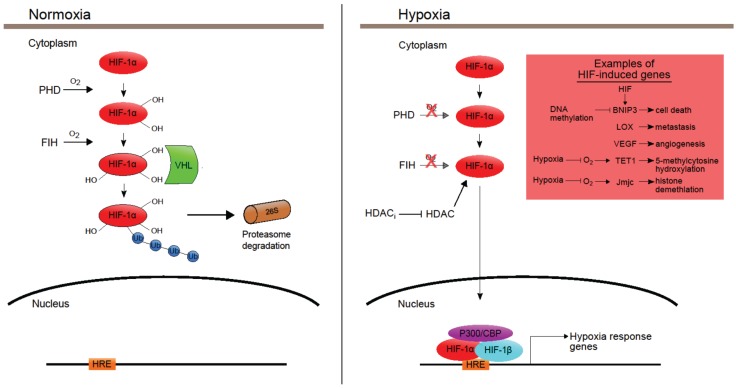
Figure [4]
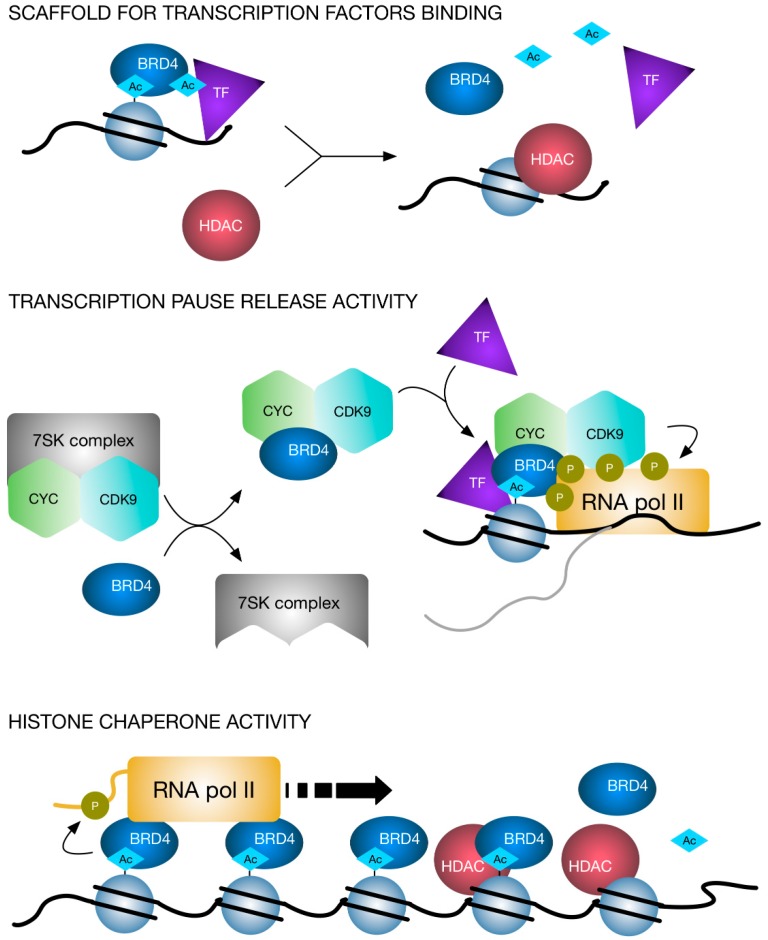
Figure [5]
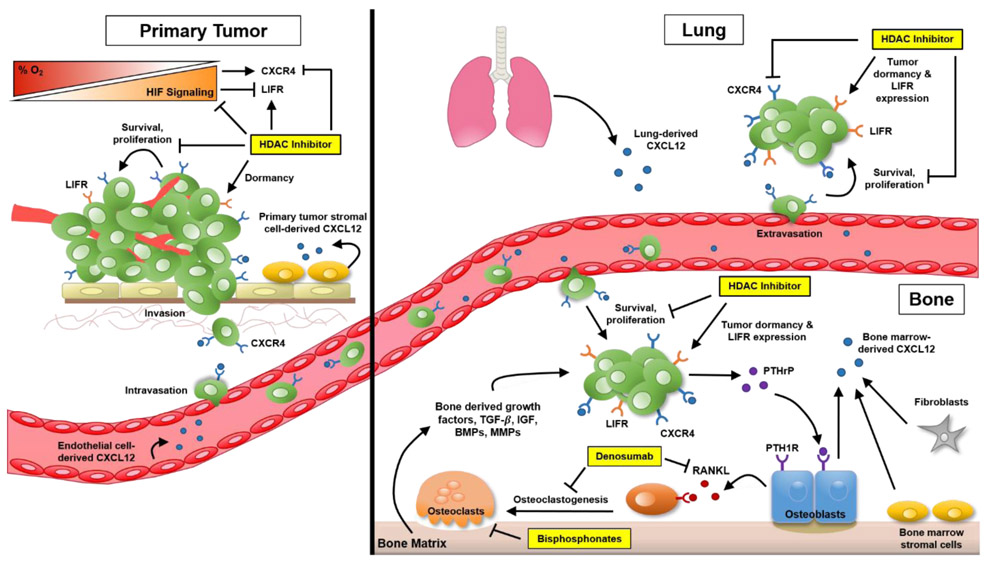
Figure [6]
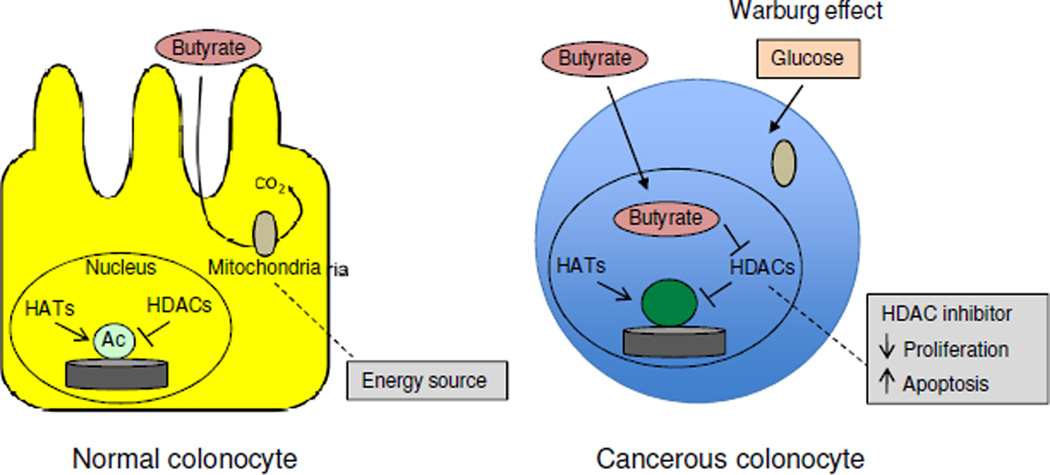
Figure [7]
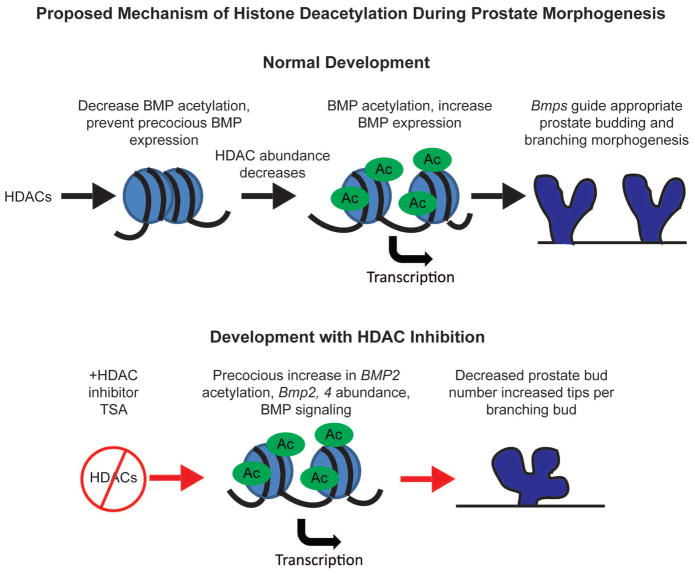
Figure [8]

Figure [9]

Figure [10]
Note: If you are interested in the full version of this target analysis report, or if you'd like to learn how our AI-powered BDE-Chem can design therapeutic molecules to interact with the HDAC9 target at a cost 90% lower than traditional approaches, please feel free to contact us at BD@silexon.ai.
More Common Targets
ABCB1 | ABCG2 | ACE2 | AHR | AKT1 | ALK | AR | ATM | BAX | BCL2 | BCL2L1 | BECN1 | BRAF | BRCA1 | CAMP | CASP3 | CASP9 | CCL5 | CCND1 | CD274 | CD4 | CD8A | CDH1 | CDKN1A | CDKN2A | CREB1 | CXCL8 | CXCR4 | DNMT1 | EGF | EGFR | EP300 | ERBB2 | EREG | ESR1 | EZH2 | FN1 | FOXO3 | HDAC9 | HGF | HMGB1 | HSP90AA1 | HSPA4 | HSPA5 | IDO1 | IFNA1 | IGF1 | IGF1R | IL17A | IL6 | INS | JUN | KRAS | MAPK1 | MAPK14 | MAPK3 | MAPK8 | MAPT | MCL1 | MDM2 | MET | MMP9 | MTOR | MYC | NFE2L2 | NLRP3 | NOTCH1 | PARP1 | PCNA | PDCD1 | PLK1 | PRKAA1 | PRKAA2 | PTEN | PTGS2 | PTK2 | RELA | SIRT1 | SLTM | SMAD4 | SOD1 | SQSTM1 | SRC | STAT1 | STAT3 | STAT5A | TAK1 | TERT | TLR4 | TNF | TP53 | TXN | VEGFA | YAP1

