As one of the major product clusters on the BDE.top platform, BDE-Chem automatically extracts and analyzes massive global drug patent / literature data, quickly screens potential active compounds based on deep learning model, accurately predicts their pharmaceutical properties and ADMET attributes, automatically completes interaction analysis by predicting the combined conformation of molecules and targets, and accelerates the whole process of drug development.
BDE-Chem
MOLECULAR GENERATION AND OPTIMIZATION
Product Introduction
By structure-activity relationship analysis, all the structure-activity relationship data from patents and literature were gathered and extracted to help researchers quickly highlight molecules, break through patents and accelerate drug discovery.
First in Class (FIC): Virtual screening of the proprietary library to generate hit and Lead compound molecules.
Fast Follow on: Through patent automatic analysis research, structure activity relationship (SAR) information extraction, and virtual screening and design, we quickly follow up on certain drugs or candidate drugs with market potential (including newly launched new drugs and candidate drugs in clinical research but with high market opportunities) and avoid these drug patents.
Repurposing: analyze existing, failed or natural products to improve active ingredients, or develop new dosage forms, new processes, new ways, new Polypill, or discover new indications.
Drug Likeness: Based on ADMET and other new energy analysis, determine the possibility of a finished product.
Structure-Activity Data Analysis
A patent analysis report can be obtained with one click, and the analysis results can be provided to predict compounds that may be possible to break through the patent, by analysis of the mother nucleus, group, similarity and side chain from tens of thousands of patents, based on the patent number entered by the user.
Molecular Recommendations
Molecular
Prediction
CPI Prediction
Based on a deep learning model, the BDE-Chem affinity module can accurately predict the interaction and affinity data between targets (proteins) and molecules (compounds), namely CPI, and quickly screen out potential active compounds for a given target in a large-scale molecular library. Based on a large-scale active database, the model has better prediction accuracy and generalization performance, and can be specifically used for data collection and model tuning for targets to achieve better performance.
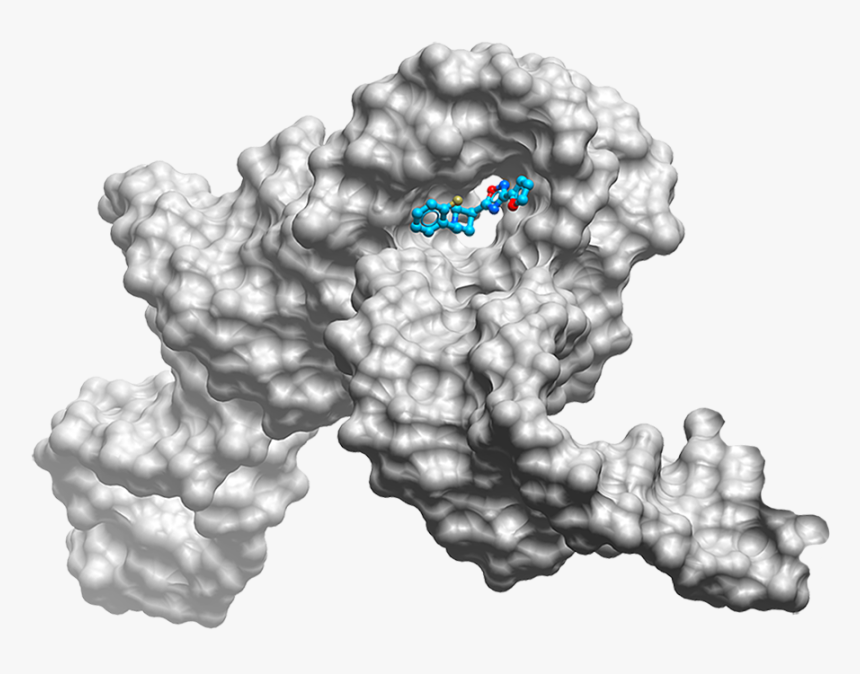
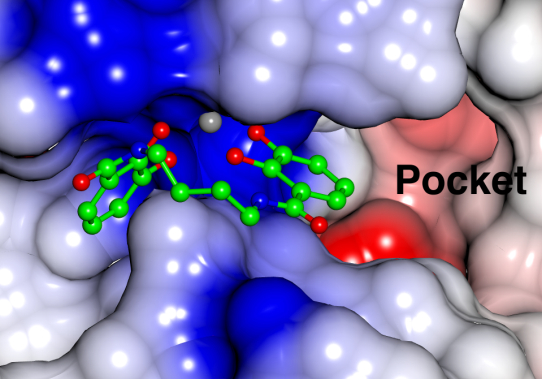
3D Combined Confirmation Prediction
Based on the Silexon's proprietory model, BDE-Chem conformation prediction module can accurately predict the binding mode of drug molecules and targets, analyze the interaction between drug molecules and targets, and provide reliable support for drug design based on three-dimensional target structure.
ADMET Prediction
Based on unique data assets and comprehensive algorithmic models, the ADMET module of the BDE-Chem provides rich ADMET attribute prediction services and AI model construction functions that combine user private data. Starting from the work scenario of a pharmaceutical chemist, data is collected and built based on actual needs, and the optimal model is iteratively optimized through continuous data updates.
Molecular Generation and Optimization
Automatic molecular generation is carried out based on the target structure following pharmacochemical constraints. The BDE-Chem molecular generation module provides a large list of potential molecules with superior comprehensive performance under the constraints of affinity, binding conformation, ADMET and other attributes, laying the foundation for subsequent molecular design and verification.
Molecules for Common Targets
ABCB1 | ABCG2 | ACE2 | AHR | AKT1 | ALK | AR | ATM | BAX | BCL2 | BCL2L1 | BRAF | BRCA1 | CASP3 | CASP9 | CCL5 | CCND1 | CD274 | CD4 | CDKN1A | CXCL8 | CXCR4 | DNMT1 | EGF | EGFR | EP300 | ERBB2 | ESR1 | EZH2 | FN1 | FOXO3 | HDAC9 | HGF | HMGB1 | HSP90AA1 | HSPA5 | IDO1 | IGF1 | IGF1R | IL17A | IL6 | INS | JUN | KRAS | MAPK1 | MAPK14 | MAPK3 | MAPK8 | MAPT | MCL1 | MMP9 | MTOR | MYC | NFE2L2 | NLRP3 | NOTCH1 | PARP1 | PCNA | PDCD1 | PLK1 | PRKAA1 | PRKAA2 | PTEN | PTGS2 | PTK2 | RELA | SIRT1 | SRC | STAT1 | STAT3 | STAT5A | TERT | TLR4 | TNF | TP53 | TXN | VEGFA | YAP1